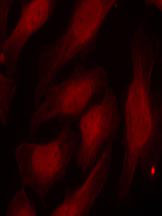
Phospho-Histone H2A.X-S139 Antibody
Purified Rabbit Polyclonal Antibody (Pab)
- SPECIFICATION
- CITATIONS: 1
- PROTOCOLS
- BACKGROUND

Application
| WB, IF |
|---|---|
| Primary Accession | P16104 |
| Reactivity | Human |
| Host | Rabbit |
| Clonality | Polyclonal |
| Concentration | 1mg/ml |
| Isotype | Rabbit IgG |
| Calculated MW | 15145 Da |
| Gene ID | 3014 |
|---|---|
| Other Names | Histone H2AX, H2a/x, Histone H2AX, H2AFX, H2AX |
| Target/Specificity | The antibody was affinity-purified from rabbit antiserum using epitope-specific phosphopeptide column, and the antibody against non-phosphopeptide was removed using non-phosphopeptide column corresponding to the phosphorylation site. |
| Dilution | WB~~1:500~1:1000 IF~~1:100~200 |
| Format | affinity Purified IgG, in PBS, 0.02% sodium azide and 50% glycerol. |
| Storage | Maintain refrigerated at 2-8°C for up to 6 months. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | Phospho-Histone H2A.X-S139 Antibody is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | H2AX (HGNC:4739) |
|---|---|
| Function | Variant histone H2A which replaces conventional H2A in a subset of nucleosomes. Nucleosomes wrap and compact DNA into chromatin, limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones thereby play a central role in transcription regulation, DNA repair, DNA replication and chromosomal stability. DNA accessibility is regulated via a complex set of post- translational modifications of histones, also called histone code, and nucleosome remodeling. Required for checkpoint-mediated arrest of cell cycle progression in response to low doses of ionizing radiation and for efficient repair of DNA double strand breaks (DSBs) specifically when modified by C-terminal phosphorylation. |
| Cellular Location | Nucleus. Chromosome |
Provided below are standard protocols that you may find useful for product applications.
Background
Histones are basic nuclear proteins that are responsible for the nucleosome structure of the chromosomal fiber in eukaryotes. Two molecules of each of the four core histones (H2A, H2B, H3, and H4) form an octamer, around which approximately 146 bp of DNA is wrapped in repeating units, called nucleosomes. The linker histone, H1, interacts with linker DNA between nucleosomes and functions in the compaction of chromatin into higher order structures. This gene encodes a member of the histone H2A family, and generates two transcripts through the use of the conserved stem-loop termination motif, and the polyA addition motif.
References
Differences in the kinetics of gamma-H2AX fluorescence decay after exposure to low and high LET radiation. Schmid TE, et al. Int J Radiat Biol, 2010 Aug. PMID 20569192.
Acetylation of H2AX on lysine 36 plays a key role in the DNA double-strand break repair pathway. Jiang X, et al. FEBS Lett, 2010 Jul 2. PMID 20488183.
H2AX phosphorylation screen of cells from radiosensitive cancer patients reveals a novel DNA double-strand break repair cellular phenotype. Vasireddy RS, et al. Br J Cancer, 2010 May 11. PMID 20461094.
High-resolution profiling of gammaH2AX around DNA double strand breaks in the mammalian genome. Iacovoni JS, et al. EMBO J, 2010 Apr 21. PMID 20360682.
Phosphorylation of histone H2A.X by DNA-dependent protein kinase is not affected by core histone acetylation, but it alters nucleosome stability and histone H1 binding. Li A, et al. J Biol Chem, 2010 Jun 4. PMID 20356835.
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.





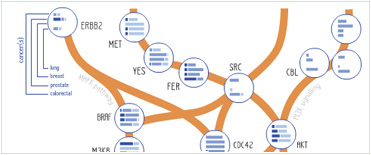







 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.