RALBP1 Antibody (Center) Blocking peptide
Synthetic peptide
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND
| Primary Accession | Q15311 |
|---|---|
| Clone Names | 101008019 |
| Gene ID | 10928 |
|---|---|
| Other Names | RalA-binding protein 1, RalBP1, 76 kDa Ral-interacting protein, Dinitrophenyl S-glutathione ATPase, DNP-SG ATPase, Ral-interacting protein 1, RALBP1, RLIP1, RLIP76 |
| Format | Peptides are lyophilized in a solid powder format. Peptides can be reconstituted in solution using the appropriate buffer as needed. |
| Storage | Maintain refrigerated at 2-8°C for up to 6 months. For long term storage store at -20°C. |
| Precautions | This product is for research use only. Not for use in diagnostic or therapeutic procedures. |
| Name | RALBP1 (HGNC:9841) |
|---|---|
| Function | Multifunctional protein that functions as a downstream effector of RALA and RALB (PubMed:7673236). As a GTPase-activating protein/GAP can inactivate CDC42 and RAC1 by stimulating their GTPase activity (PubMed:7673236). As part of the Ral signaling pathway, may also regulate ligand-dependent EGF and insulin receptors-mediated endocytosis (PubMed:10910768, PubMed:12775724). During mitosis, may act as a scaffold protein in the phosphorylation of EPSIN/EPN1 by the mitotic kinase cyclin B-CDK1, preventing endocytosis during that phase of the cell cycle (PubMed:12775724). During mitosis, also controls mitochondrial fission as an effector of RALA (PubMed:21822277). Recruited to mitochondrion by RALA, acts as a scaffold to foster the mitotic kinase cyclin B-CDK1-mediated phosphorylation and activation of DNM1L (PubMed:21822277). |
| Cellular Location | Cell membrane; Peripheral membrane protein. Cytoplasm, cytosol Cytoplasm, cytoskeleton, spindle pole {ECO:0000250|UniProtKB:Q62796} Nucleus. Mitochondrion. Note=Cytosolic protein that transiently associates with the mitotic spindle poles in early prophase, and dissociates from them after completion of mitosis (By similarity) Targeted to the plasma membrane through its interaction with RALB, directed by FGF signaling. Docking on the membrane is required to transduce the Ral signal (By similarity). Recruited by RALA to the mitochondrion during mitosis where it regulates mitochondrial fission (PubMed:21822277). Nuclear localization is cell cycle dependent while membrane localization is seen in adherent cells (PubMed:22319010). The region involved in membrane association could form transmembrane domains and expose a part of the protein extracellularly (Probable) {ECO:0000250|UniProtKB:Q62796, ECO:0000250|UniProtKB:Q9PT60, ECO:0000269|PubMed:21822277, ECO:0000269|PubMed:22319010, ECO:0000305|PubMed:15610018} |
| Tissue Location | Expressed ubiquitously but at low levels. Shows a strong expression in the erythrocytes. |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
Background
RALBP1 plays a role in receptor-mediated endocytosis andis a downstream effector of the small GTP-binding protein RAL (seeRALA; MIM 179550). Small G proteins, such as RAL, have GDP-boundinactive and GTP-bound active forms, which shift from the inactiveto the active state through the action of RALGDS (MIM 601619),which in turn is activated by RAS (see HRAS; MIM 190020) (summaryby Feig, 2003 [PubMed 12888294]).
References
Singhal, S.S., et al. Int. J. Cancer 126(6):1327-1338(2010)Singhal, S.S., et al. Cancer Lett. 283(2):152-158(2009)Awasthi, Y.C., et al. J Toxicol Environ Health B Crit Rev 12(7):540-551(2009)Singhal, S.S., et al. Cancer Res. 69(10):4244-4251(2009)Singhal, S.S., et al. Biochem. Pharmacol. 77(6):1074-1083(2009)
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.




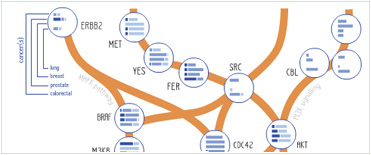







 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.