SUMOplot™ Analysis Program
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. The presence of this post-translational modification may help explain larger MWs than expected on SDS gels due to attachment of SUMO protein (11kDa) at multiple positions in your protein.
SUMO-1 (small ubiquitin-related modifier; also known as PIC1, UBL1, Sentrin, GMP1, and Smt3) is a member of the ubiquitin and ubiquitin-like superfamily. Most SUMO-modified proteins contain the tetrapeptide motif B-K-x-D/E where B is a hydrophobic residue, K is the lysine conjugated to SUMO, x is any amino acid (aa), D or E is an acidic residue.
The SUMOplot™ Analysis Program predicts the probability for the SUMO consensus sequence (SUMO-CS) to be engaged in SUMO attachment. The SUMOplot™ score system is based on two criteria: direct amino acid match to SUMO-CS. substitution of the consensus amino acid residues with amino acid residues exhibiting similar hydrophobicity.
Developed by Abcepta, Copyright © 2013 -2025
|
|





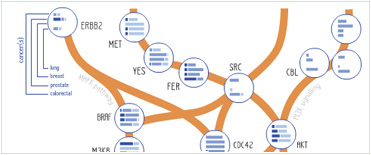







 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.