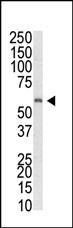
MBD1 Antibody (C-term)
Purified Rabbit Polyclonal Antibody (Pab)
- SPECIFICATION
- CITATIONS: 2
- PROTOCOLS
- BACKGROUND

Application
| WB, E |
|---|---|
| Primary Accession | Q9UIS9 |
| Reactivity | Human |
| Host | Rabbit |
| Clonality | Polyclonal |
| Isotype | Rabbit IgG |
| Calculated MW | 66607 Da |
| Antigen Region | 376-405 aa |
| Gene ID | 4152 |
|---|---|
| Other Names | Methyl-CpG-binding domain protein 1, CXXC-type zinc finger protein 3, Methyl-CpG-binding protein MBD1, Protein containing methyl-CpG-binding domain 1, MBD1, CXXC3, PCM1 |
| Target/Specificity | This MBD1 antibody is generated from rabbits immunized with a KLH conjugated synthetic peptide between 376-405 amino acids from the C-terminal region of human MBD1. |
| Dilution | WB~~1:1000 E~~Use at an assay dependent concentration. |
| Format | Purified polyclonal antibody supplied in PBS with 0.09% (W/V) sodium azide. This antibody is prepared by Saturated Ammonium Sulfate (SAS) precipitation followed by dialysis against PBS. |
| Storage | Maintain refrigerated at 2-8°C for up to 2 weeks. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | MBD1 Antibody (C-term) is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | MBD1 (HGNC:6916) |
|---|---|
| Synonyms | CXXC3, PCM1 |
| Function | Transcriptional repressor that binds CpG islands in promoters where the DNA is methylated at position 5 of cytosine within CpG dinucleotides. Binding is abolished by the presence of 7-mG that is produced by DNA damage by methylmethanesulfonate (MMS). Acts as transcriptional repressor and plays a role in gene silencing by recruiting ATF7IP, which in turn recruits factors such as the histone methyltransferase SETDB1. Probably forms a complex with SETDB1 and ATF7IP that represses transcription and couples DNA methylation and histone 'Lys-9' trimethylation. Isoform 1 and isoform 2 can also repress transcription from unmethylated promoters. |
| Cellular Location | Nucleus. Nucleus matrix. Nucleus speckle Chromosome Note=Nuclear, in a punctate pattern (PubMed:12711603). Associated with euchromatic regions of the chromosomes, with pericentromeric regions on chromosome 1 and with telomeric regions from several chromosomes (PubMed:10454587, PubMed:10648624). |
| Tissue Location | Widely expressed.. |
Provided below are standard protocols that you may find useful for product applications.
Background
DNA methylation, or the addition of methyl groups to cytosine bases in the dinucleotide CpG, is imperative to proper development and regulates gene expression. The methylation pattern involves the enzymatic processes of methylation and demethylation. The demethylation enzyme was recently found to be a mammalian protein, which exhibits demethylase activity associated to a methyl-CpG-binding domain (MBD) (1). The enzyme is able to revert methylated cytosine bases to cytosines within the particular dinucleotide sequence mdCpdG by catalyzing the cleaving of the methyl group as methanol. MeCP2 and MBD1 (PCM1) are first found to repress transcription by binding specifically to methylated DNA (2). MBD2 and MBD4 (also known as MED1) were later found to colocalize with foci of heavily methylated satellite DNA and believed to mediate the biological functions of the methylation signal. Surprisingly, MBD3 does not bind methylated DNA both in vivo and in vitro. MBD1, MBD2, MBD3, and MBD4 are found to be expressed in somatic tissues, but the expression of MBD1 and MBD2 is reduced or absent in embryonic stem cells, which are known to be deficient in MeCP1 activity. MBD4 have homology to bacterial base excision repair DNA N-glycosylases/lyases (3). In some microsatellite unstable tumors MBD4 is mutated at an exonic polynucleotide tract (4).
References
Bhattacharya SK, Ramchandani S, Cervoni N, Szyf. M. Nature, 397 (6720):579-583 1999.
Hendrich B and Bird A. Mol Cell Biol, 18: 6538-6547(1998).
Petronzelli F, Riccio A, Markham GD, Seeholzer SH, Stoerker J, Genuardi M, Yeung AT, Matsumoto Y, Bellacosa A. J Biol Chem 275 (42): 32422-32429 (2000).
Bader S, Walker M, Harrison D. Br J Cancer 83(12): 1646-1649 (2000).
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.





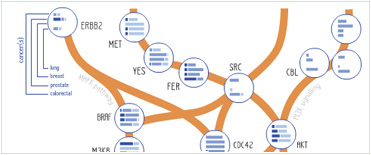







 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.

