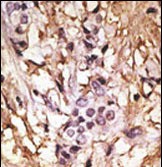
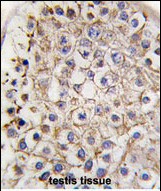
LC3 Antibody (APG8B) (C-term)
Purified Rabbit Polyclonal Antibody (Pab)
- SPECIFICATION
- CITATIONS: 13
- PROTOCOLS
- BACKGROUND

Application
| IHC-P, WB, E |
|---|---|
| Primary Accession | Q9GZQ8 |
| Other Accession | A6NCE7 |
| Reactivity | Human |
| Host | Rabbit |
| Clonality | Polyclonal |
| Isotype | Rabbit IgG |
| Calculated MW | 14688 Da |
| Antigen Region | 77-106 aa |
| Gene ID | 81631 |
|---|---|
| Other Names | Microtubule-associated proteins 1A/1B light chain 3B, Autophagy-related protein LC3 B, Autophagy-related ubiquitin-like modifier LC3 B, MAP1 light chain 3-like protein 2, MAP1A/MAP1B light chain 3 B, MAP1A/MAP1B LC3 B, Microtubule-associated protein 1 light chain 3 beta, MAP1LC3B, MAP1ALC3 |
| Target/Specificity | This LC3 antibody is generated from rabbits immunized with a KLH conjugated synthetic peptide between 77-106 amino acids from the C-terminal region of human LC3. |
| Dilution | IHC-P~~1:50~100 WB~~1:1000 E~~Use at an assay dependent concentration. |
| Format | Purified polyclonal antibody supplied in PBS with 0.09% (W/V) sodium azide. This antibody is prepared by Saturated Ammonium Sulfate (SAS) precipitation followed by dialysis against PBS. |
| Storage | Maintain refrigerated at 2-8°C for up to 2 weeks. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | LC3 Antibody (APG8B) (C-term) is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | MAP1LC3B (HGNC:13352) |
|---|---|
| Synonyms | MAP1ALC3 |
| Function | Ubiquitin-like modifier involved in formation of autophagosomal vacuoles (autophagosomes) (PubMed:20418806, PubMed:23209295, PubMed:28017329). Plays a role in mitophagy which contributes to regulate mitochondrial quantity and quality by eliminating the mitochondria to a basal level to fulfill cellular energy requirements and preventing excess ROS production (PubMed:23209295, PubMed:28017329). In response to cellular stress and upon mitochondria fission, binds C-18 ceramides and anchors autophagolysosomes to outer mitochondrial membranes to eliminate damaged mitochondria (PubMed:22922758). While LC3s are involved in elongation of the phagophore membrane, the GABARAP/GATE-16 subfamily is essential for a later stage in autophagosome maturation (PubMed:20418806, PubMed:23209295, PubMed:28017329). Promotes primary ciliogenesis by removing OFD1 from centriolar satellites via the autophagic pathway (PubMed:24089205). Through its interaction with the reticulophagy receptor TEX264, participates in the remodeling of subdomains of the endoplasmic reticulum into autophagosomes upon nutrient stress, which then fuse with lysosomes for endoplasmic reticulum turnover (PubMed:31006537, PubMed:31006538). Upon nutrient stress, directly recruits cofactor JMY to the phagophore membrane surfaces and promotes JMY's actin nucleation activity and autophagosome biogenesis during autophagy (PubMed:30420355). |
| Cellular Location | Cytoplasmic vesicle, autophagosome membrane; Lipid-anchor Endomembrane system; Lipid-anchor Mitochondrion membrane; Lipid-anchor. Cytoplasm, cytoskeleton {ECO:0000250|UniProtKB:Q9CQV6}. Cytoplasmic vesicle. Note=LC3-II binds to the autophagic membranes. LC3-II localizes with the mitochondrial inner membrane during Parkin-mediated mitophagy (PubMed:28017329). Also localizes to discrete punctae along the ciliary axoneme |
| Tissue Location | Most abundant in heart, brain, skeletal muscle and testis. Little expression observed in liver |
Provided below are standard protocols that you may find useful for product applications.
Background
Macroautophagy is the major inducible pathway for the general turnover of cytoplasmic constituents in eukaryotic cells, it is also responsible for the degradation of active cytoplasmic enzymes and organelles during nutrient starvation. Macroautophagy involves the formation of double-membrane bound autophagosomes which enclose the cytoplasmic constituent targeted for degradation in a membrane bound structure, which then fuse with the lysosome (or vacuole) releasing a single-membrane bound autophagic bodies which are then degraded within the lysosome (or vacuole). MAP1A and MAP1B are microtubule-associated proteins which mediate the physical interactions between microtubules and components of the cytoskeleton. These proteins are involved in formation of autophagosomal vacuoles (autophagosomes). MAP1A and MAP1B each consist of a heavy chain subunit and multiple light chain subunits. MAP1LC3b is one of the light chain subunits and can associate with either MAP1A or MAP1B. The precursor molecule is cleaved by APG4B/ATG4B to form the cytosolic form, LC3-I. This is activated by APG7L/ATG7, transferred to ATG3 and conjugated to phospholipid to form the membrane-bound form, LC3-II.
References
Baehrecke EH. Nat Rev Mol Cell Biol. 6(6):505-10. (2005) Lum JJ, et al. Nat Rev Mol Cell Biol. 6(6):439-48. (2005) Greenberg JT. Dev Cell. 8(6):799-801. (2005) Levine B. Cell. 120(2):159-62. (2005) Shintani T and Klionsky DJ. Science. 306(5698):990-5. (2004) Tanida I., et al. Int. J. Biochem. Cell Biol. 36:2503-2518(2004) He H., et al. J. Biol. Chem. 278:29278-29287(2003) Tanida I., et al. J. Biol. Chem. 279:36268-36276(2004)
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.







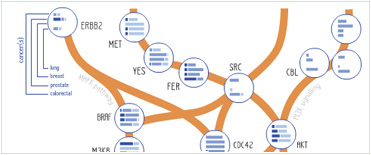







 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.