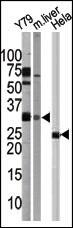
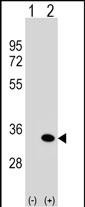
ATG5 Antibody (C-term)
Purified Rabbit Polyclonal Antibody (Pab)
- SPECIFICATION
- CITATIONS: 28
- PROTOCOLS
- BACKGROUND

Application
| IF, WB, IHC-P, E |
|---|---|
| Primary Accession | Q9H1Y0 |
| Reactivity | Human, Mouse |
| Host | Rabbit |
| Clonality | Polyclonal |
| Isotype | Rabbit IgG |
| Calculated MW | 32447 Da |
| Antigen Region | 209-238 aa |
| Gene ID | 9474 |
|---|---|
| Other Names | Autophagy protein 5, APG5-like, Apoptosis-specific protein, ATG5, APG5L, ASP |
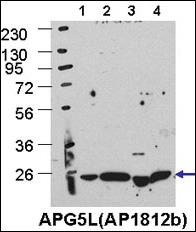
| Target/Specificity | This ATG5 antibody is generated from rabbits immunized with a KLH conjugated synthetic peptide between 209-238 amino acids from the C-terminal region of human ATG5. |
| Dilution | IF~~1:25 WB~~1:1000 IHC-P~~1:50~100 E~~Use at an assay dependent concentration. |
| Format | Purified polyclonal antibody supplied in PBS with 0.09% (W/V) sodium azide. This antibody is purified through a protein A column, followed by peptide affinity purification. |
| Storage | Maintain refrigerated at 2-8°C for up to 2 weeks. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | ATG5 Antibody (C-term) is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | ATG5 (HGNC:589) |
|---|---|
| Synonyms | APG5L, ASP |
| Function | Involved in autophagic vesicle formation. Conjugation with ATG12, through a ubiquitin-like conjugating system involving ATG7 as an E1-like activating enzyme and ATG10 as an E2-like conjugating enzyme, is essential for its function. The ATG12-ATG5 conjugate acts as an E3- like enzyme which is required for lipidation of ATG8 family proteins and their association to the vesicle membranes. Involved in mitochondrial quality control after oxidative damage, and in subsequent cellular longevity. Plays a critical role in multiple aspects of lymphocyte development and is essential for both B and T lymphocyte survival and proliferation. Required for optimal processing and presentation of antigens for MHC II. Involved in the maintenance of axon morphology and membrane structures, as well as in normal adipocyte differentiation. Promotes primary ciliogenesis through removal of OFD1 from centriolar satellites and degradation of IFT20 via the autophagic pathway. As part of the ATG8 conjugation system with ATG12 and ATG16L1, required for recruitment of LRRK2 to stressed lysosomes and induction of LRRK2 kinase activity in response to lysosomal stress (By similarity). |
| Cellular Location | Cytoplasm. Preautophagosomal structure membrane; Peripheral membrane protein. Note=Colocalizes with nonmuscle actin. The conjugate detaches from the membrane immediately before or after autophagosome formation is completed (By similarity). Also localizes to discrete punctae along the ciliary axoneme and to the base of the ciliary axoneme. Under starved conditions, the ATG12-ATG5-ATG16L1 complex is translocated to phagophores driven by RAB33B (PubMed:32960676). {ECO:0000250, ECO:0000269|PubMed:32960676} |
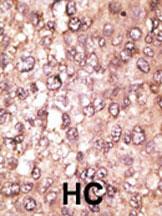
| Tissue Location | Ubiquitous. The mRNA is present at similar levels in viable and apoptotic cells, whereas the protein is dramatically highly expressed in apoptotic cells |
Provided below are standard protocols that you may find useful for product applications.
Background
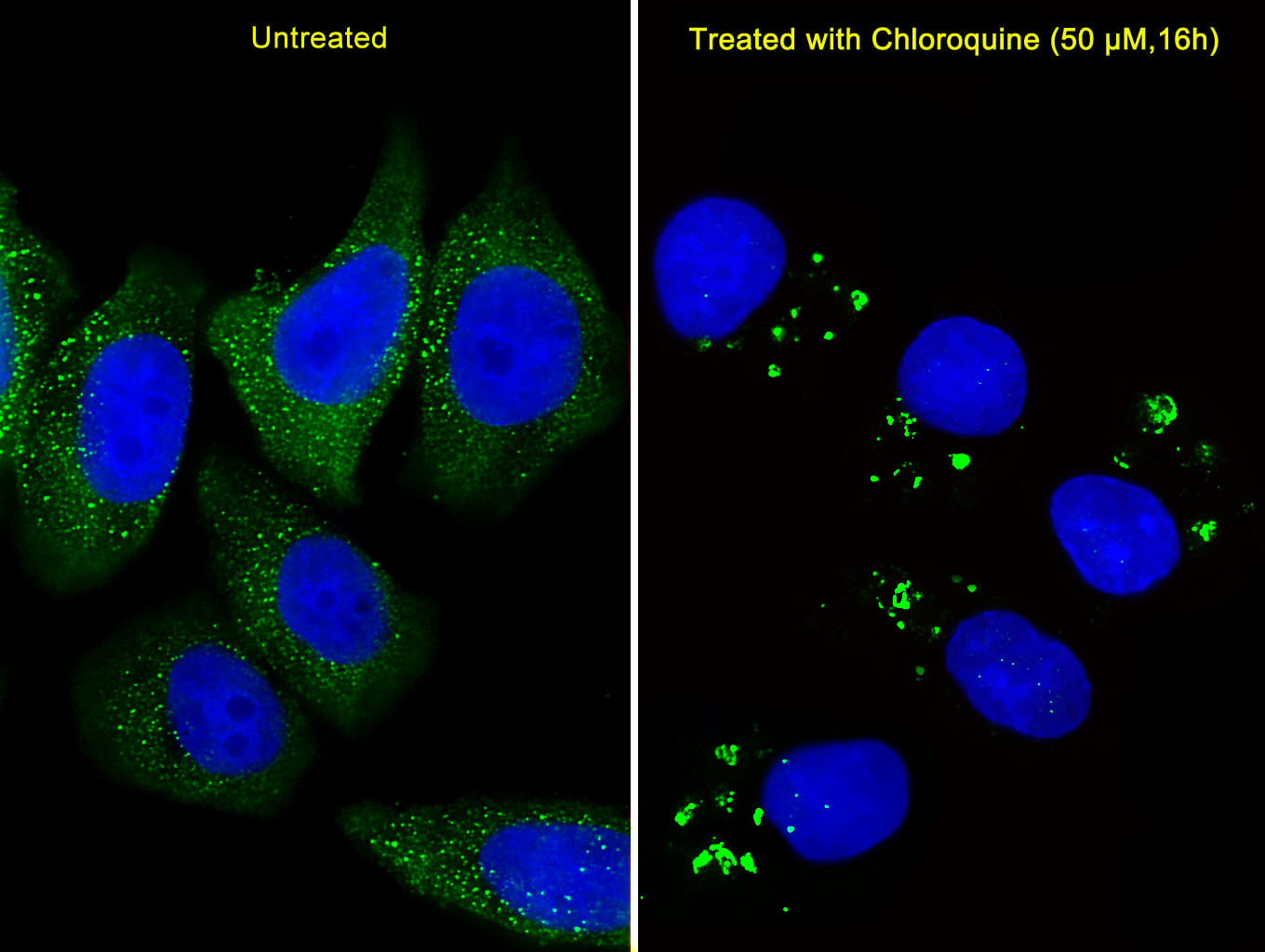
Macroautophagy is the major inducible pathway for the general turnover of cytoplasmic constituents in eukaryotic cells, it is also responsible for the degradation of active cytoplasmic enzymes and organelles during nutrient starvation. Macroautophagy involves the formation of double-membrane bound autophagosomes which enclose the cytoplasmic constituent targeted for degradation in a membrane bound structure, which then fuse with the lysosome (or vacuole) releasing a single-membrane bound autophagic bodies which are then degraded within the lysosome (or vacuole). APG5, required for autophagy, conjugates to ATG12 and associates with an isolation membrane to form a cup-shaped isolation membrane and autophagosome. The conjugate detaches from the membrane immediately before or after autophagosome formation is completed. APG5 may also play an important role in the apoptotic process, possibly within the modified cytoskeleton. Its expression is a relatively late event in the apoptotic process, occurring downstream of caspase activity.
References
Baehrecke EH. Nat Rev Mol Cell Biol. 6(6):505-10. (2005) Lum JJ, et al. Nat Rev Mol Cell Biol. 6(6):439-48. (2005) Greenberg JT. Dev Cell. 8(6):799-801. (2005) Levine B. Cell. 120(2):159-62. (2005) Shintani T and Klionsky DJ. Science. 306(5698):990-5. (2004) Hammond E.M., et al. FEBS Lett. 425:391-395(1998) Strausberg R.L., et al. PNAS 99:16899-16903(2002) Grand R.J.A., et al. Exp. Cell Res. 218:439-451(1995) Mizushima N., et al. J. Biol. Chem. 273:33889-33892(1998) Mizushima N., et al. J. Cell Biol. 152:657-668(2001)
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.





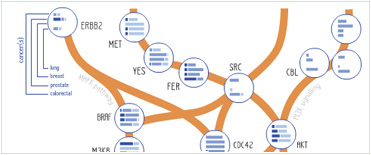







 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.