ARFBP1 Polyclonal Antibody
Purified Rabbit Polyclonal Antibody (Pab)
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
| IHC-P, IHC-F, IF, ICC, E |
|---|---|
| Primary Accession | Q7Z6Z7 |
| Reactivity | Rat, Pig, Dog, Bovine |
| Host | Rabbit |
| Clonality | Polyclonal |
| Calculated MW | 482 KDa |
| Physical State | Liquid |
| Immunogen | KLH conjugated synthetic peptide derived from human ARF6 |
| Epitope Specificity | 3201-3400/43741 |
| Isotype | IgG |
| Purity | affinity purified by Protein A |
| Buffer | 0.01M TBS (pH7.4) with 1% BSA, 0.02% Proclin300 and 50% Glycerol. |
| SUBCELLULAR LOCATION | Cytoplasm. Nucleus. Mainly expressed in the cytoplasm of most tissues, except in the nucleus of spermatogonia, primary spermatocytes and neuronal cells (By similarity). Predominantly cytosolic or perinuclear in some colorectal carcinoma cells. |
| SIMILARITY | Belongs to the TOM1/PTR1 family. Contains 1 HECT (E6AP-type E3 ubiquitin-protein ligase) domain. Contains 1 UBA domain. Contains 1 UIM (ubiquitin-interacting motif) repeat. Contains 1 WWE domain. |
| SUBUNIT | Interacts with isoform p14ARF of CDKN2A which strongly inhibits HUWE1 ubiquitin ligase activity. Interacts with MYCN, POLB and CDC6. |
| Post-translational modifications | Phosphorylated on tyrosine; phosphorylation is probably required for its ability to inhibit TP53 transactivation. Phosphorylated upon DNA damage, probably by ATM or ATR. |
| DISEASE | Defects in HUWE1 are the cause of mental retardation syndromic X-linked Turner type (MRXST) [MIM:300706]; also known as mental retardation and macrocephaly syndrome. MRXST shows clinical variability. Associated phenotypes include macrocephaly and variable contractures. A chromosomal microduplication involving HUWE1 and HSD17B10 is the cause of mental retardation X-linked type 17 (MRX17) [MIM:300705]; also known as mental retardation X-linked type 31 (MRX31). Mental retardation is characterized by significantly sub-average general intellectual functioning associated with impairments in adaptative behavior and manifested during the developmental period. In contrast to syndromic or specific X-linked mental retardation which also present with associated physical, neurological and/or psychiatric manifestations, intellectual deficiency is the only primary symptom of non-syndromic X-linked mental retardation. |
| Important Note | This product as supplied is intended for research use only, not for use in human, therapeutic or diagnostic applications. |
| Background Descriptions | E3 ubiquitin-protein ligase which mediates ubiquitination and subsequent proteasomal degradation of target proteins. Regulates apoptosis by catalyzing the polyubiquitination and degradation of MCL1. Mediates monoubiquitination of DNA polymerase beta (POLB) at 'Lys-41', 'Lys-61' and 'Lys-81', thereby playing a role in base-excision repair. Also ubiquitinates the p53/TP53 tumor suppressor and core histones including H1, H2A, H2B, H3 and H4. Binds to an upstream initiator-like sequence in the preprodynorphin gene. Regulates neural differentiation and proliferation by catalyzing the polyubiquitination and degradation of MYCN. May regulate abundance of CDC6 after DNA damage by polyubiquitinating and targeting CDC6 to degradation. |
| Gene ID | 10075 |
|---|---|
| Other Names | E3 ubiquitin-protein ligase HUWE1, 2.3.2.26, ARF-binding protein 1, ARF-BP1, HECT, UBA and WWE domain-containing protein 1, HECT-type E3 ubiquitin transferase HUWE1, Homologous to E6AP carboxyl terminus homologous protein 9, HectH9, Large structure of UREB1, LASU1, Mcl-1 ubiquitin ligase E3, Mule, Upstream regulatory element-binding protein 1, URE-B1, URE-binding protein 1, HUWE1, KIAA0312, KIAA1578, UREB1 |
| Target/Specificity | Weakly expressed in heart, brain and placenta but not in other tissues. Expressed in a number of cell lines, predominantly in those from colorectal carcinomas. |
| Dilution | IHC-P=1:100-500,IHC-F=1:100-500,ICC=1:100-500,IF=1:100-500,ELISA=1:5000-10000 |
| Storage | Store at -20 ℃ for one year. Avoid repeated freeze/thaw cycles. When reconstituted in sterile pH 7.4 0.01M PBS or diluent of antibody the antibody is stable for at least two weeks at 2-4 ℃. |
| Name | HUWE1 |
|---|---|
| Synonyms | KIAA0312, KIAA1578, UREB1 |
| Function | E3 ubiquitin-protein ligase which mediates ubiquitination and subsequent proteasomal degradation of target proteins (PubMed:15567145, PubMed:15767685, PubMed:15989957, PubMed:17567951, PubMed:18488021, PubMed:19037095, PubMed:19713937, PubMed:20534529, PubMed:30217973). Regulates apoptosis by catalyzing the polyubiquitination and degradation of MCL1 (PubMed:15989957). Mediates monoubiquitination of DNA polymerase beta (POLB) at 'Lys-41', 'Lys-61' and 'Lys-81', thereby playing a role in base-excision repair (PubMed:19713937). Also ubiquitinates the p53/TP53 tumor suppressor and core histones including H1, H2A, H2B, H3 and H4 (PubMed:15567145, PubMed:15767685, PubMed:15989956). Ubiquitinates MFN2 to negatively regulate mitochondrial fusion in response to decreased stearoylation of TFRC (PubMed:26214738). Ubiquitination of MFN2 also takes place following induction of mitophagy; AMBRA1 acts as a cofactor for HUWE1-mediated ubiquitination (PubMed:30217973). Regulates neural differentiation and proliferation by catalyzing the polyubiquitination and degradation of MYCN (PubMed:18488021). May regulate abundance of CDC6 after DNA damage by polyubiquitinating and targeting CDC6 to degradation (PubMed:17567951). Mediates polyubiquitination of isoform 2 of PA2G4 (PubMed:19037095). Acts in concert with MYCBP2 to regulate the circadian clock gene expression by promoting the lithium-induced ubiquination and degradation of NR1D1 (PubMed:20534529). Binds to an upstream initiator-like sequence in the preprodynorphin gene (By similarity). Mediates HAPSTR1 degradation, but is also a required cofactor in the pathway by which HAPSTR1 governs stress signaling (PubMed:35776542). Acts as a regulator of the JNK and NF-kappa-B signaling pathways by mediating assembly of heterotypic 'Lys-63'-/'Lys- 48'-linked branched ubiquitin chains that are then recognized by TAB2: HUWE1 mediates branching of 'Lys-48'-linked chains of substrates initially modified with 'Lys-63'-linked conjugates by TRAF6 (PubMed:27746020). 'Lys-63'-/'Lys-48'-linked branched ubiquitin chains protect 'Lys-63'-linkages from CYLD deubiquitination (PubMed:27746020). Ubiquitinates PPARA in hepatocytes (By similarity). |
| Cellular Location | Cytoplasm. Nucleus. Mitochondrion. Note=Mainly expressed in the cytoplasm of most tissues, except in the nucleus of spermatogonia, primary spermatocytes and neuronal cells (By similarity). Recruited to mitochondria following interaction with AMBRA1 (PubMed:30217973) {ECO:0000250|UniProtKB:Q7TMY8, ECO:0000269|PubMed:30217973} |
| Tissue Location | Weakly expressed in heart, brain and placenta but not in other tissues. Expressed in a number of cell lines, predominantly in those from colorectal carcinomas |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.




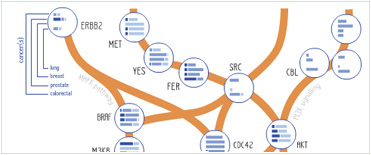







 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.