NFKBIL2 Polyclonal Antibody
Purified Rabbit Polyclonal Antibody (Pab)
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
| IHC-P, IHC-F, IF, ICC, E |
|---|---|
| Primary Accession | Q96HA7 |
| Reactivity | Rat, Dog |
| Host | Rabbit |
| Clonality | Polyclonal |
| Calculated MW | 150929 Da |
| Gene ID | 4796 |
|---|---|
| Other Names | Tonsoku-like protein, Inhibitor of kappa B-related protein, I-kappa-B-related protein, IkappaBR, NF-kappa-B inhibitor-like protein 2, Nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor-like 2, TONSL, IKBR, NFKBIL2 |
| Dilution | IHC-P=1:100-500,IHC-F=1:100-500,ICC=1:100-500,IF=1:100-500,ELISA=1:5000-10000 |
| Storage | Store at -20 ℃ for one year. Avoid repeated freeze/thaw cycles. When reconstituted in sterile pH 7.4 0.01M PBS or diluent of antibody the antibody is stable for at least two weeks at 2-4 ℃. |
| Name | TONSL {ECO:0000303|PubMed:21055983, ECO:0000312|HGNC:HGNC:7801} |
|---|---|
| Function | Component of the MMS22L-TONSL complex, a complex that promotes homologous recombination-mediated repair of double-strand breaks (DSBs) at stalled or collapsed replication forks (PubMed:21055983, PubMed:21055984, PubMed:21055985, PubMed:21113133, PubMed:26527279, PubMed:27338793, PubMed:27797818, PubMed:29478807, PubMed:30773278). The MMS22L-TONSL complex is required to maintain genome integrity during DNA replication (PubMed:21055983, PubMed:21055984, PubMed:21055985). It mediates the assembly of RAD51 filaments on single-stranded DNA (ssDNA): the MMS22L-TONSL complex is recruited to DSBs following histone replacement by histone chaperones and eviction of the replication protein A complex (RPA/RP-A) from DSBs (PubMed:21055983, PubMed:21055984, PubMed:21055985, PubMed:27797818, PubMed:29478807). Following recruitment to DSBs, the TONSL-MMS22L complex promotes recruitment of RAD51 filaments and subsequent homologous recombination (PubMed:27797818, PubMed:29478807). Within the complex, TONSL acts as a histone reader, which recognizes and binds newly synthesized histones following their replacement by histone chaperones (PubMed:27338793, PubMed:29478807). Specifically binds histone H4 lacking methylation at 'Lys-20' (H4K20me0) and histone H3.1 (PubMed:27338793). |
| Cellular Location | Nucleus. Chromosome. Cytoplasm Note=Mainly nuclear (PubMed:21055983, PubMed:21055984). Localizes to DNA damage sites, accumulates at stressed replication forks (PubMed:21055983, PubMed:21055984, PubMed:26527279, PubMed:27338793) Recruited to stalled or collapsed replication forks following histone replacement by histone chaperones ASF1A and the CAF-1 complex: TONSL acts as a histone reader that recognizes and binds newly synthesized histones (PubMed:29478807). |
| Tissue Location | Expressed in heart, skeletal muscle and tracheal epithelial cells. |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.




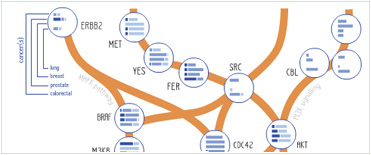







 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.