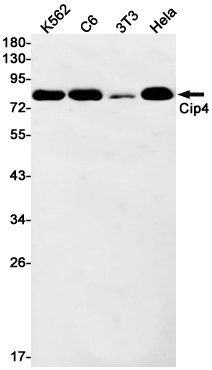
CIP4 Rabbit mAb
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
| WB |
|---|---|
| Primary Accession | Q15642 |
| Reactivity | Human, Mouse, Rat |
| Host | Rabbit |
| Clonality | Monoclonal Antibody |
| Calculated MW | 68352 Da |
| Gene ID | 9322 |
|---|---|
| Other Names | TRIP10 |
| Dilution | WB~~1/500-1/1000 |
| Format | 50mM Tris-Glycine(pH 7.4), 0.15M NaCl, 40%Glycerol, 0.01% sodium azide and 0.05% BSA. |
| Storage | Store at 4°C short term. Aliquot and store at -20°C long term. Avoid freeze/thaw cycles. |
| Name | TRIP10 |
|---|---|
| Synonyms | CIP4, STOT, STP |
| Function | Required for translocation of GLUT4 to the plasma membrane in response to insulin signaling (By similarity). Required to coordinate membrane tubulation with reorganization of the actin cytoskeleton during endocytosis. Binds to lipids such as phosphatidylinositol 4,5- bisphosphate and phosphatidylserine and promotes membrane invagination and the formation of tubules. Also promotes CDC42-induced actin polymerization by recruiting WASL/N-WASP which in turn activates the Arp2/3 complex. Actin polymerization may promote the fission of membrane tubules to form endocytic vesicles. Required for the formation of podosomes, actin-rich adhesion structures specific to monocyte- derived cells. May be required for the lysosomal retention of FASLG/FASL. |
| Cellular Location | Cytoplasm, cytoskeleton. Cytoplasm, cell cortex. Lysosome. Golgi apparatus. Cell membrane. Cell projection, phagocytic cup. Note=Translocates to the plasma membrane in response to insulin stimulation, and this may require active RHOQ (By similarity) Localizes to cortical regions coincident with F-actin, to lysosomes and to sites of phagocytosis in macrophages. Also localizes to the Golgi, and this requires AKAP9. |
| Tissue Location | Expressed in brain, colon, heart, kidney, liver, lung, megakaryocyte, ovary, pancreas, peripheral blood lymphocytes, placenta, prostate, skeletal muscle, small intestine, spleen, testis, thymus and trachea. |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.




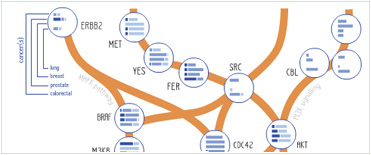







 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.