SETDB1 Antibody (C-term) Blocking Peptide
Synthetic peptide
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND
| Primary Accession | Q15047 |
|---|---|
| Clone Names | 4020606 |
| Gene ID | 9869 |
|---|---|
| Other Names | Histone-lysine N-methyltransferase SETDB1, ERG-associated protein with SET domain, ESET, Histone H3-K9 methyltransferase 4, H3-K9-HMTase 4, Lysine N-methyltransferase 1E, SET domain bifurcated 1, SETDB1, KIAA0067, KMT1E |
| Target/Specificity | The synthetic peptide sequence used to generate the antibody AP1073b was selected from the C-term region of human SETDB1. A 10 to 100 fold molar excess to antibody is recommended. Precise conditions should be optimized for a particular assay. |
| Format | Peptides are lyophilized in a solid powder format. Peptides can be reconstituted in solution using the appropriate buffer as needed. |
| Storage | Maintain refrigerated at 2-8°C for up to 6 months. For long term storage store at -20°C. |
| Precautions | This product is for research use only. Not for use in diagnostic or therapeutic procedures. |
| Name | SETDB1 (HGNC:10761) |
|---|---|
| Function | Histone methyltransferase that specifically trimethylates 'Lys-9' of histone H3. H3 'Lys-9' trimethylation represents a specific tag for epigenetic transcriptional repression by recruiting HP1 (CBX1, CBX3 and/or CBX5) proteins to methylated histones. Mainly functions in euchromatin regions, thereby playing a central role in the silencing of euchromatic genes. H3 'Lys-9' trimethylation is coordinated with DNA methylation (PubMed:12869583). Required for HUSH-mediated heterochromatin formation and gene silencing. Forms a complex with MBD1 and ATF7IP that represses transcription and couples DNA methylation and histone 'Lys-9' trimethylation (PubMed:27732843, PubMed:14536086). Its activity is dependent on MBD1 and is heritably maintained through DNA replication by being recruited by CAF-1 (PubMed:14536086). SETDB1 is targeted to histone H3 by TRIM28/TIF1B, a factor recruited by KRAB zinc-finger proteins. Probably forms a corepressor complex required for activated KRAS-mediated promoter hypermethylation and transcriptional silencing of tumor suppressor genes (TSGs) or other tumor-related genes in colorectal cancer (CRC) cells (PubMed:24623306). Required to maintain a transcriptionally repressive state of genes in undifferentiated embryonic stem cells (ESCs) (PubMed:24623306). In ESCs, in collaboration with TRIM28, is also required for H3K9me3 and silencing of endogenous and introduced retroviruses in a DNA- methylation independent-pathway (By similarity). Associates at promoter regions of tumor suppressor genes (TSGs) leading to their gene silencing (PubMed:24623306). The SETDB1-TRIM28-ZNF274 complex may play a role in recruiting ATRX to the 3'-exons of zinc-finger coding genes with atypical chromatin signatures to establish or maintain/protect H3K9me3 at these transcriptionally active regions (PubMed:27029610). |
| Cellular Location | Nucleus. Cytoplasm. Chromosome. Note=Associated with non- pericentromeric regions of chromatin. Excluded from nucleoli and islands of condensed chromatin. |
| Tissue Location | Widely expressed. High expression in testis. |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
Background
The SET domain is a highly conserved, approximately 150-amino acid motif implicated in the modulation of chromatin structure. It was originally identified as part of a larger conserved region present in the Drosophila Trithorax protein and was subsequently identified in the Drosophila Su(var)3-9 and 'Enhancer of zeste' proteins, from which the acronym SET is derived. Studies have suggested that the SET domain may be a signature of proteins that modulate transcriptionally active or repressed chromatin states through chromatin remodeling activities.
References
Ichimura, T., et al., J. Biol. Chem. 280(14):13928-13935 (2005).Sarraf, S.A., et al., Mol. Cell 15(4):595-605 (2004).Wang, H., et al., Mol. Cell 12(2):475-487 (2003).Schultz, D.C., et al., Genes Dev. 16(8):919-932 (2002).Yang, L., et al., Biochem. J. 369 (PT 3), 651-657 (2003) (): ().
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.




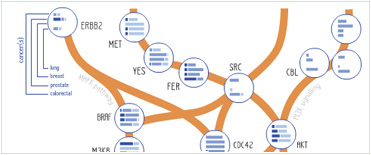







 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.