USP10 Antibody (N-term) Blocking Peptide
Synthetic peptide
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND
| Primary Accession | Q14694 |
|---|---|
| Clone Names | 3063004 |
| Gene ID | 9100 |
|---|---|
| Other Names | Ubiquitin carboxyl-terminal hydrolase 10, Deubiquitinating enzyme 10, Ubiquitin thioesterase 10, Ubiquitin-specific-processing protease 10, USP10, KIAA0190 |
| Target/Specificity | The synthetic peptide sequence used to generate the antibody AP2138a was selected from the N-term region of human USP10 . A 10 to 100 fold molar excess to antibody is recommended. Precise conditions should be optimized for a particular assay. |
| Format | Peptides are lyophilized in a solid powder format. Peptides can be reconstituted in solution using the appropriate buffer as needed. |
| Storage | Maintain refrigerated at 2-8°C for up to 6 months. For long term storage store at -20°C. |
| Precautions | This product is for research use only. Not for use in diagnostic or therapeutic procedures. |
| Name | USP10 {ECO:0000303|PubMed:11439350, ECO:0000312|HGNC:HGNC:12608} |
|---|---|
| Function | Hydrolase that can remove conjugated ubiquitin from target proteins such as p53/TP53, RPS2/us5, RPS3/us3, RPS10/eS10, BECN1, SNX3 and CFTR (PubMed:11439350, PubMed:18632802, PubMed:31981475). Acts as an essential regulator of p53/TP53 stability: in unstressed cells, specifically deubiquitinates p53/TP53 in the cytoplasm, leading to counteract MDM2 action and stabilize p53/TP53 (PubMed:20096447). Following DNA damage, translocates to the nucleus and deubiquitinates p53/TP53, leading to regulate the p53/TP53-dependent DNA damage response (PubMed:20096447). Component of a regulatory loop that controls autophagy and p53/TP53 levels: mediates deubiquitination of BECN1, a key regulator of autophagy, leading to stabilize the PIK3C3/VPS34-containing complexes (PubMed:21962518). In turn, PIK3C3/VPS34-containing complexes regulate USP10 stability, suggesting the existence of a regulatory system by which PIK3C3/VPS34-containing complexes regulate p53/TP53 protein levels via USP10 and USP13 (PubMed:21962518). Does not deubiquitinate MDM2 (PubMed:20096447). Plays a key role in 40S ribosome subunit recycling when a ribosome has stalled during translation: acts both by inhibiting formation of stress granules, which store stalled translation pre-initiation complexes, and mediating deubiquitination of 40S ribosome subunits (PubMed:27022092, PubMed:31981475, PubMed:34348161, PubMed:34469731). Acts as a negative regulator of stress granules formation by lowering G3BP1 and G3BP2 valence, thereby preventing G3BP1 and G3BP2 ability to undergo liquid- liquid phase separation (LLPS) and assembly of stress granules (PubMed:11439350, PubMed:27022092, PubMed:32302570). Promotes 40S ribosome subunit recycling following ribosome dissociation in response to ribosome stalling by mediating deubiquitination of 40S ribosomal proteins RPS2/us5, RPS3/us3 and RPS10/eS10, thereby preventing their degradation by the proteasome (PubMed:31981475, PubMed:34348161, PubMed:34469731). Part of a ribosome quality control that takes place when ribosomes have stalled during translation initiation (iRQC): USP10 acts by removing monoubiquitination of RPS2/us5 and RPS3/us3, promoting 40S ribosomal subunit recycling (PubMed:34469731). Deubiquitinates CFTR in early endosomes, enhancing its endocytic recycling (PubMed:19398555). Involved in a TANK-dependent negative feedback response to attenuate NF-kappa-B activation via deubiquitinating IKBKG or TRAF6 in response to interleukin-1-beta (IL1B) stimulation or upon DNA damage (PubMed:25861989). Deubiquitinates TBX21 leading to its stabilization (PubMed:24845384). Plays a negative role in the RLR signaling pathway upon RNA virus infection by blocking the RIGI- mediated MAVS activation. Mechanistically, removes the unanchored 'Lys- 63'-linked polyubiquitin chains of MAVS to inhibit its aggregation, essential for its activation (PubMed:37582970). |
| Cellular Location | Cytoplasm. Nucleus. Early endosome. Note=Cytoplasmic in normal conditions (PubMed:20096447). After DNA damage, translocates to the nucleus following phosphorylation by ATM (PubMed:20096447) |
| Tissue Location | Widely expressed.. |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
Background
Modification of target proteins by ubiquitin participates in a wide array of biological functions. Proteins destined for degradation or processing via the 26 S proteasome are coupled to multiple copies of ubiquitin. However, attachment of ubiquitin or ubiquitin-related molecules may also result in changes in subcellular distribution or modification of protein activity. An additional level of ubiquitin regulation, deubiquitination, is catalyzed by proteases called deubiquitinating enzymes, which fall into four distinct families. Ubiquitin C-terminal hydrolases, ubiquitin-specific processing proteases (USPs),1 OTU-domain ubiquitin-aldehyde-binding proteins, and Jab1/Pad1/MPN-domain-containing metallo-enzymes. Among these four families, USPs represent the most widespread and represented deubiquitinating enzymes across evolution. USPs tend to release ubiquitin from a conjugated protein. They display similar catalytic domains containing conserved Cys and His boxes but divergent N-terminal and occasionally C-terminal extensions, which are thought to function in substrate recognition, subcellular localization, and protein-protein interactions.
References
Strausberg, R.L., et al., Proc. Natl. Acad. Sci. U.S.A. 99(26):16899-16903 (2002).Soncini, C., et al., Oncogene 20(29):3869-3879 (2001).Nagase, T., et al., DNA Res. 3(1):17-24 (1996).
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.




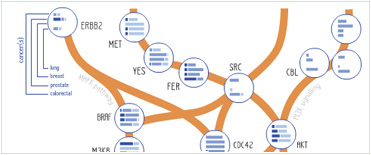







 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.