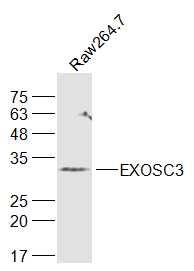
EXOSC3 Polyclonal Antibody
Purified Rabbit Polyclonal Antibody (Pab)
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
| WB, IHC-P, IHC-F, IF, ICC, E |
|---|---|
| Primary Accession | Q9NQT5 |
| Reactivity | Rat |
| Host | Rabbit |
| Clonality | Polyclonal |
| Calculated MW | 29 KDa |
| Physical State | Liquid |
| Immunogen | KLH conjugated synthetic peptide derived from human EXOSC3 |
| Epitope Specificity | 101-200/275 |
| Isotype | IgG |
| Purity | affinity purified by Protein A |
| Buffer | 0.01M TBS (pH7.4) with 1% BSA, 0.02% Proclin300 and 50% Glycerol. |
| SUBCELLULAR LOCATION | Cytoplasm. Nucleus, nucleolus. |
| SIMILARITY | Belongs to the RRP40 family. |
| SUBUNIT | Component of the RNA exosome complex. Specifically part of the catalytically inactive RNA exosome core (Exo-9) complex which is believed to associate with catalytic subunits EXOSC10, and DIS3 or DIS3L in cytoplasmic- and nuclear-specific RNA exosome complex forms. Exo-9 is formed by a hexameric ring of RNase PH domain-containing subunits specifically containing the heterodimers EXOSC4-EXOSC9, EXOSC5-EXOSC8 and EXOSC6-EXOSC7, and peripheral S1 domain-containing components EXOSC1, EXOSC2 and EXOSC3 located on the top of the ring structure. Interacts with GTPBP1. Interacts with ZC3HAV1. Interacts with DDX17 only in the presence of ZC3HAV1 in an RNA-independent manner. |
| DISEASE | Defects in EXOSC3 are the cause of pontocerebellar hypoplasia type 1B (PCH1B) [MIM:614678]. A severe autosomal recessive neurologic disorder characterized by a combination of cerebellar and spinal motor neuron degeneration beginning at birth. There is diffuse muscle weakness, progressive microcephaly, global developmental delay, and brainstem involvement. |
| Important Note | This product as supplied is intended for research use only, not for use in human, therapeutic or diagnostic applications. |
| Background Descriptions | The exosome is a multisubunit complex of 3’ to 5’ exoribonucleases. It is involved in a variety of cellular processes and is responsible for degrading unstable mRNAs that contain AU-rich elements in their untranslated 3’ region. EXOSC3 (exosome component 3), also known as p10, CGI-102, RRP40 (Ribosomal RNA-processing protein 40), Rrp40p or hRrp40p, is a component of the exosome multienzyme ribonuclease complex. Localizing to the cytoplasm and nucleolus, EXOSC3 contains a putative S1 RNA-binding domain and is capable of binding RNA. EXOSC3 is a component of the top cap of the exosome and is essential for exosome stability. In addition, EXOSC3 is required for the processing of the 7S pre-RNA to the mature 5.8S rRNA. |
| Gene ID | 51010 |
|---|---|
| Other Names | Exosome complex component RRP40, Exosome component 3, Ribosomal RNA-processing protein 40, p10, EXOSC3, RRP40 |
| Dilution | WB=1:500-2000,IHC-P=1:100-500,IHC-F=1:100-500,ICC=1:100-500,IF=1:100-500,ELISA=1:5000-10000 |
| Storage | Store at -20 ℃ for one year. Avoid repeated freeze/thaw cycles. When reconstituted in sterile pH 7.4 0.01M PBS or diluent of antibody the antibody is stable for at least two weeks at 2-4 ℃. |
| Name | EXOSC3 |
|---|---|
| Synonyms | RRP40 |
| Function | Non-catalytic component of the RNA exosome complex which has 3'->5' exoribonuclease activity and participates in a multitude of cellular RNA processing and degradation events. In the nucleus, the RNA exosome complex is involved in proper maturation of stable RNA species such as rRNA, snRNA and snoRNA, in the elimination of RNA processing by-products and non-coding 'pervasive' transcripts, such as antisense RNA species and promoter-upstream transcripts (PROMPTs), and of mRNAs with processing defects, thereby limiting or excluding their export to the cytoplasm. The RNA exosome may be involved in Ig class switch recombination (CSR) and/or Ig variable region somatic hypermutation (SHM) by targeting AICDA deamination activity to transcribed dsDNA substrates. In the cytoplasm, the RNA exosome complex is involved in general mRNA turnover and specifically degrades inherently unstable mRNAs containing AU-rich elements (AREs) within their 3' untranslated regions, and in RNA surveillance pathways, preventing translation of aberrant mRNAs. It seems to be involved in degradation of histone mRNA. The catalytic inactive RNA exosome core complex of 9 subunits (Exo-9) is proposed to play a pivotal role in the binding and presentation of RNA for ribonucleolysis, and to serve as a scaffold for the association with catalytic subunits and accessory proteins or complexes. EXOSC3 as peripheral part of the Exo-9 complex stabilizes the hexameric ring of RNase PH-domain subunits through contacts with EXOSC9 and EXOSC5. |
| Cellular Location | Cytoplasm. Nucleus, nucleolus. Nucleus |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.




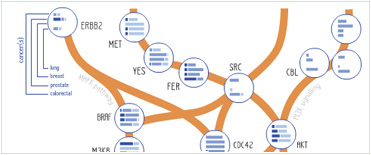







 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.